
APA Style
Sudheer Kumar Katari, Chandu Sai Kanakala, Anil Kumar Singh. (2026). Computational Exploration of Mutant SARS-CoV-2 Main Protease as a Target for Binding and Inhibition Attributes Utilizing Antiviral Drugs for Therapeutic Application . Molecular Modeling Connect, 3 (Article ID: 0016). https://doi.org/Registering DOIMLA Style
Sudheer Kumar Katari, Chandu Sai Kanakala, Anil Kumar Singh. "Computational Exploration of Mutant SARS-CoV-2 Main Protease as a Target for Binding and Inhibition Attributes Utilizing Antiviral Drugs for Therapeutic Application ". Molecular Modeling Connect, vol. 3, 2026, Article ID: 0016, https://doi.org/Registering DOI.Chicago Style
Sudheer Kumar Katari, Chandu Sai Kanakala, Anil Kumar Singh. 2026. "Computational Exploration of Mutant SARS-CoV-2 Main Protease as a Target for Binding and Inhibition Attributes Utilizing Antiviral Drugs for Therapeutic Application ." Molecular Modeling Connect 3 (2026): 0016. https://doi.org/Registering DOI.
 ACCESS
Research Article
ACCESS
Research Article
Volume 3, Article ID: 2026.0016

Sudheer Kumar Katari
katari319@gmail.com

Chandu Sai Kanakala
chandukanakala811@gmail.com

Anil Kumar Singh
phd.anil@yahoo.com
1 Department of Biotechnology, Vignan’s Foundation for Science, Technology and Research, Vadlamudi-522213, Guntur, India
2 Academy of Scientific and Innovative Research (AcSIR), Ghaziabad-201002, India.
* Author to whom correspondence should be addressed
Received: 02 Nov 2025 Accepted: 20 Mar 2026 Available Online: 17 Apr 2026
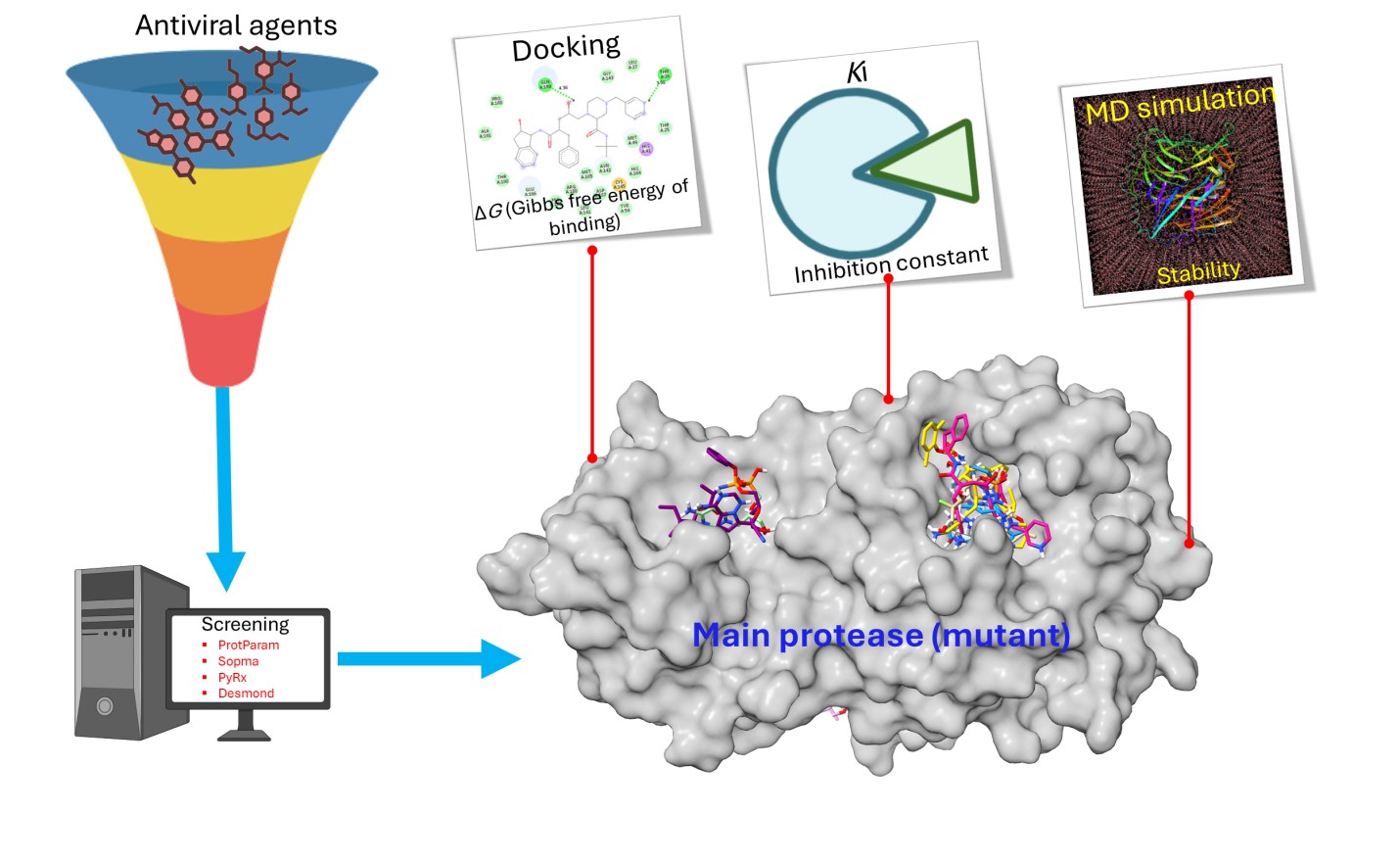
Mutations in the main protease (Mpro) of SARS-CoV-2 have resulted in resistance to available inhibitors and various other compounds that have been reported. Targeting the inhibition of SARS-CoV-2 Mpro presents a promising opportunity to develop potent therapeutic agents aimed at combating SARS-CoV-2 and similar viruses. The presented study evaluates a selection of ten antiviral agents—Acyclovir, Remdesivir, Sofosbuvir, Atazanavir, Cidofovir, Indinavir, Lopinavir, Oseltamivir, Galidesivir, and Favipiravir—focusing on their ability to bind and inhibit mutant Mpro through various computational methods. The computational physical-chemical properties and secondary structure elements (SSE) exhibit comparable values between WT and mutant variants of Mpro. The Mpro-Indinavir docked complex was identified as the high-ranked docked compound, with a binding affinity of -8.07±0.06 Kcal/mol and a Ki value of 1.21 µM. This docked complex was involved hydrogen bond interactions between the THR-26 and GLN-189 residues. The binding affinity of all docked compounds was found to be in a range from -5.37± 0.40 to -8.07±0.06 Kcal/mol. The docked complexes underwent a thorough assessment through a 100 ns MD simulation, which confirmed the stability of the top-ranked docked complexes. The findings indicate that some antiviral agents exhibit adequate binding affinity and inhibitory effectiveness, as predicted by Ki. However, theoretical findings necessitate the subsequent experimental validation. This suggests that compounds with notable binding and other properties can be effectively repurposed as promising therapeutic candidates for Mpro inhibition targeting mutant variants.
Disclaimer: This is not the final version of the article. Changes may occur when the manuscript is published in its final format.
We use cookies to improve your experience on our site. By continuing to use our site, you accept our use of cookies. Learn more